Research
Immune-mediated diseases, their relationships and divisions
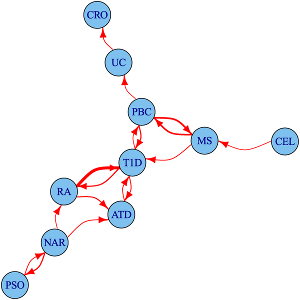
We study the shared and distinct genetic aetiology between related autoimmune diseases both at the genomewide and region specific level, addressing two separate questions. First, at a the level of the whole genome, do associations with one disease predict associations with another disease, and if so, can we exploit these to improve genetic discovery? We do this using approaches which control the conditional false discovery rate of a statistic which combines the observed p values with informative data about each genetic variant, often the p values from a related disease. More recently, we developed a functional approach to this problem which is faster and more powerful.
Because this uses genomewide associations, it includes information from individual loci that are not genomewide significant, but show a trend to shared association for two diseases. However, because genes with similar function are often located close to one another, and because genetic variants also show a spatial correlation due to linkage disequilibrium, this does not autoimattically imply that the two diseases share the same causal variants.

To address this more specific question, we use a complementary approach called colocalisation that matches two GWAS signatures in a local region to investigate sharing of causal variants in detail. We maintain the coloc package for colocalisation, and have led its development in areas such as allowing multiple causal variants through integration with the SuSiE method for fine-mapping, and incorporating variant-specific priors.
Mapping causal genes
Genomewide association studies (GWAS) have been hugely successful in identifying associations between genetic variation and risk of common diseases. However, we need to translate this knowledge into understanding the genes, cells and pathways involved. This is a difficult problem because the associated genetic variants do not typically reside in genes and change the protein they encode, but lie between genes and are presumed to regulate their expression in some cells, perhaps under specific conditions.
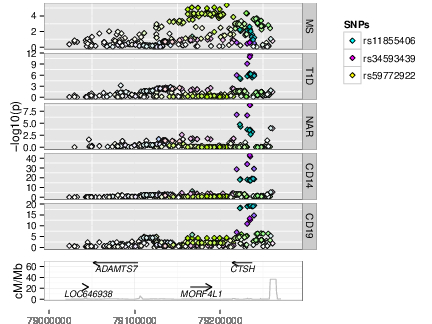
As GWAS studies for disease have been conducted over the past decade, so people have conducted GWAS for other, more gene specific traits, such as parallel GWAS for the expression levels of each of the ~20,000 protein coding genes in a given cell type, called “eQTL” studies. These produce a local GWAS trace just as a disease GWAS does. We can use colocalisation to determine whether the two GWAS traces are compatible with the two traits, disease risk and expression of a given gene in a given cell type and condition, sharing a causal variant.
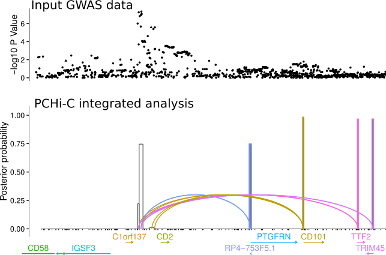
Another approach is to exploit knowledge about the 3D folding of chromatin derived from high throughput Chromosome Conformation Capture (Hi-C) in its targetted form: Capture Hi-C (CHi-C). This allows us to link GWAS causal variants (mapped probabilistically) to the genes they regulate, and we have deployed this approach across 17 primary human sorted cell types, as well as a separate more detailed comparison using CD4+ T cells, both activated and non-activated.
Gene expression in immune cells
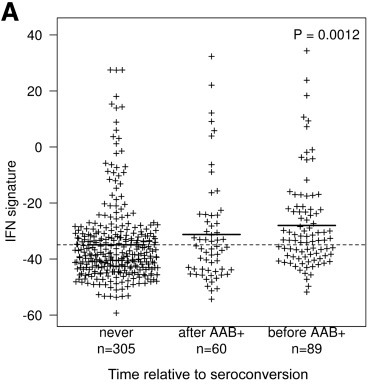
The BABYDIET study led by Annette Ziegler and Ezio Bonifacio collected blood samples longitudinally from 109 children genetically at risk of, but initially unaffected by type 1 diabetes. We were lucky to have access to white blood cells (PBMCs) from these samples and measured the gene expression in them. We were particularly interested in expression of interferon responsive genes, given previous links between type 1 diabetes and infection. Indeed, we saw an upregulation of these genes in children who went on to develop the autoantibodies that are strongly predictive of T1D diagnosis, but, crucially, the upregulation was transient and preceded the appearance of autoantibodies. It was also temporally correlated with recent upper respiratory infection, and may represent a biomarker for the response to infection or the mechanism by which the infection influences type 1 diabetes risk.
We also used these data to investigate seasonal variation in gene expression, and found that 25% of the genes expressed in these cells in these children varied in their expression throughout the year. In winter, we saw that this expression profile produced a pro-inflammatory environment. This might be advantageous during a season when infectious diseases are at a peak, but is a risk factor for other diseases associated with inflammation such as cardiovascular also peak in winter. We replicated this finding in multiple datasets, including one from the Southern hemisphere when winter occurs during June-August.
We have also developed methods to cluster samples according to gene expression signatures and to simultaneously cluster genes and samples to learn new signatures.
